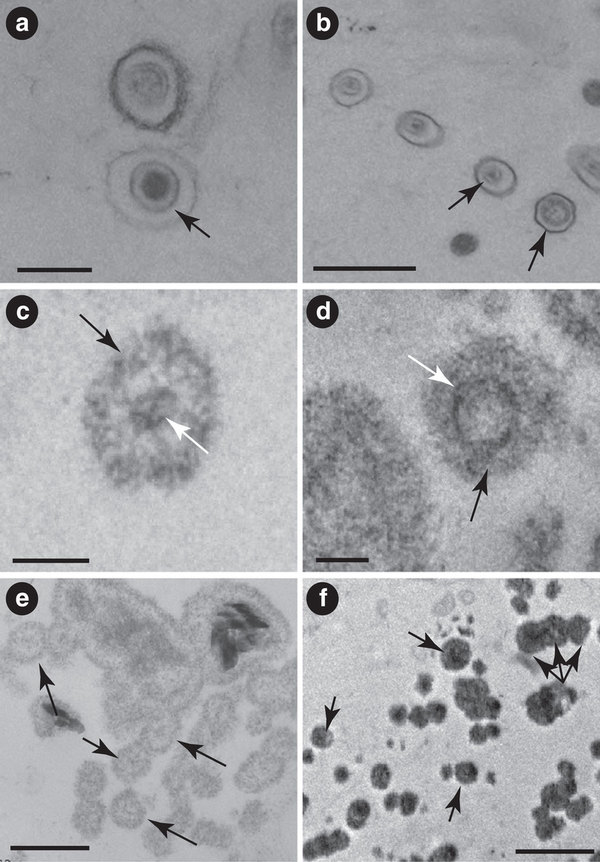
A guest post by Dr. Paul Hyman,
Ashland University, Ashland, OH, USA
phyman@ashland.edu
Bacteriophage capsids have been studied as frameworks for the development of new materials. In part this is an outgrowth of phage display which allows the precise placement of binding elements on the phage surface. But another approach is to use the native capsids and to nonspecifically attach conductive metals to create wire-like structures or to capture and encapsulate molecules in the capsids as the capsids assemble. In a recent paper in Nature Chemistry, Trevor Douglas’ group at Indiana University and colleagues have shown that the efficiency of an enzyme that reduces protons to form hydrogen gas is more efficient and stable when enclosed in a bacteriophage capsid.
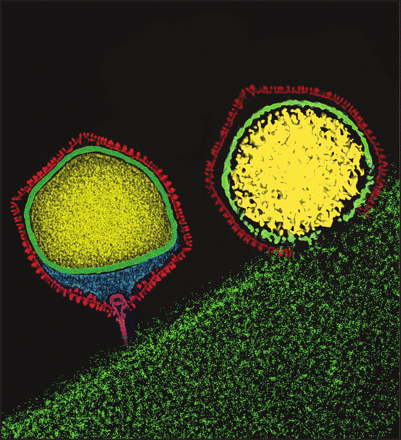
The bacteriophage they used is the Salmonella phage P22. P22’s capsid is an icosahedron composed of a major coat protein assembled onto a scaffold protein. Coat protein and scaffold protein self-assemble into a procapsid that during a normal infection is then packaged with the phage DNA. The researchers fused the two subunits of a hydrogenase protein to separate scaffold protein genes. When expressed together with coat protein, procapsid self-assembled with the heterodimeric hydrogenase protein inside as outlined in this figure.

(modified from Jordan 2015)
When they tested for hydrogenase activity, the highest efficiency was found if the scaffold protein/hydrogenase subunits were expressed several hours before the coat protein (line one in the table). This pre-encapsulation period presumably allowed the two hydrogenase subunits to assemble into the active heterodimer before being constrained by the coat protein.
|
pH 5 |
pH 8 |
| P22 encapsulated with sequential expression |
6118 nmol H2/mg min |
3218 nmol H2/mg min |
| P22 encapsulated with simultaneous expression |
|
757 nmol H2/mg min |
| Unencapsulated hydrogenase + scaffold without coat protein |
46 nmol H2/mg min |
12.6 nmol H2/mg min |
| Free hydrogenase |
12-38 nmol H2/mg min (pH not specified) |
Data from Jordan 2015
Additional experiments showed that the encapsulation also partially protected the hydrogenase against trypsin, heat denaturation (60°C for 45 min.) and air exposure.
The reason for the increased enzymatic activity is not entirely clear. The increased activity and protection results suggest that the enzyme’s quaternary structure is stabilized in some way in the capsid. It may also be that enzyme efficiency is higher when several hundred copies of the enzyme are in close proximity to each other in some sort of synergistic effect.
Overall, this paper demonstrates another way that phages can be used in non-biological technologies as well as biological. Independent of the phage aspect, an improved catalyst for production of hydrogen gas could prove quite valuable as alternative fuels, such as hydrogen, are increasingly sought after.
Reference: Paul C. Jordan, Dustin P. Patterson, Kendall N. Saboda, Ethan J. Edwards, Heini M. Miettinen, Gautam Basu, Megan C. Thielges, and Trevor Douglas, “Self-Assembling Biomolecular Catalysts for Hydrogen Production”, Nature Chemistry doi:10.1038/nchem.2416, published on-line December 21, 2015.



 “From a modern microbiological viewpoint the motivation behind the experiments that led to the discovery of bacteriophage is hard to understand. What was the purpose of filtering a bacterial culture to remove the bacteria, then remixing the filtrate with a fresh bacterial culture? We all know the outcome of this classic experiment, but why was it done in the first place?
“From a modern microbiological viewpoint the motivation behind the experiments that led to the discovery of bacteriophage is hard to understand. What was the purpose of filtering a bacterial culture to remove the bacteria, then remixing the filtrate with a fresh bacterial culture? We all know the outcome of this classic experiment, but why was it done in the first place?